Overview
The HyperANF (Hyper-Approximate Neighborhood Function) algorithm is used to evaluate the average graph distance. It offers a trade-off between accuracy and computational efficiency, making it suitable for large-scale graphs where computing the exact average distance may be infeasible or time-consuming.
Related material of the algorithm:
- P. Boldi, M. Rosa, S. Vigna, HyperANF: Approximating the Neighbourhood Function of Very Large Graphs on a Budget (2011)
Concepts
Average Graph Distance
The average graph distance is a metric used to measure the average number of steps or edges required to traverse between any two nodes in a graph. It quantifies the overall connectivity or closeness of the nodes in the graph.
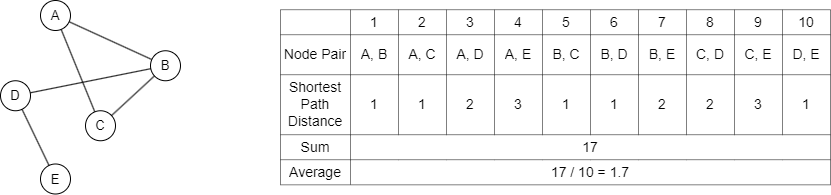
As is shown above, the average graph distance is typically computed by performing a graph traversal to calculate the shortest path distance between every pair of nodes, then summing up the distances and dividing by the total number of node pairs to get the average.
Approximate Neighborhood Function (ANF)
Graph traversals can be computationally expensive and memory-intensive, especially for large-scale graphs. In such cases, approximate neighborhood function (ANF) algorithms are commonly used to estimate the average graph distance more efficiently.
ANF algorithms aim to estimate the neighborhood function (NF):
- The neighborhood function (NF) of a graph, denoted as
N(t), returns the number of node pairs such that the two nodes can reach each other with t or fewer steps. - The individual neighborhood function (INF) of a node x in a graph, denoted as
N(x,t), returns the number of nodes that can be reached from x with t or fewer steps. - In an undirect graph G = (V, E), the relationship between NF and INF is:
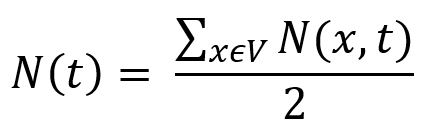
The NF can help to reveal some features of graphs, including the average graph distance:
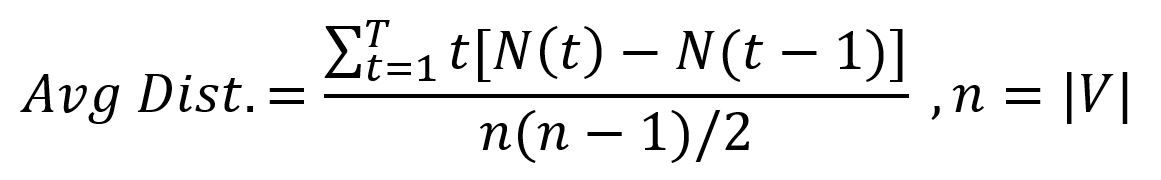
The calculation of the above example graph is shown below:

However, it is very expensive to compute the NF exactly on large graphs. By approximating the neighborhood function, ANF algorithms can estimate the average graph distance without traversing the entire graph.
HyperLogLog Counter
HyperLogLog counter is used to count approximately the number of distinct elements (i.e., the cardinality) in a large set or stream of elements. Calculating the exact cardinality often requires an amount of memory proportional to the cardinality, which is impractical for very large data sets. HyperLoglog takes a significantly less memory, with the space complexity as O(log(log n)) (this is the reason why these counters are called HyperLogLog).
A HyperLogLog counter can be viewed as an array of m = 2b registers, and each register is initialized to -∞. For example, b = 3, then M[0] = M[1] = ... = M[7] = -∞.
The number of registers depends on the desired precision of the estimation. More registers can provide a more accurate estimation, but also require more memory.
First, each element x in the set is mapped into a fixed-size binary sequence by a hash function h(). For example, h(x) = 0100001110101....
Then, update the registers. For each element x in the set:
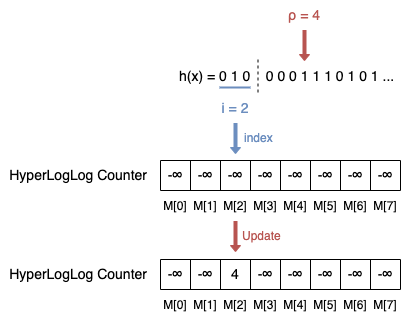
- Calculate the index i of the register by the integer value of the leftmost b bits of h(x), i.e., hb(x). In the example, i = hb(x) = 010 = 0*22 + 1*21 + 0*20 = 2.
- Let hb(x) be the sequence of remaining bits of h(x), and ρ(hb(x)) be the position of the leftmost 1 of hb(x). In the example, ρ(hb(x)) = ρ(0001110101...) = 4.
- Update register M[i] = max(M[i], ρ(hb(x))). In the example, M[2] = max(-∞, 4) = 4.
After reading all elements, the cardinality is calculated by the HyperLogLog counter as:
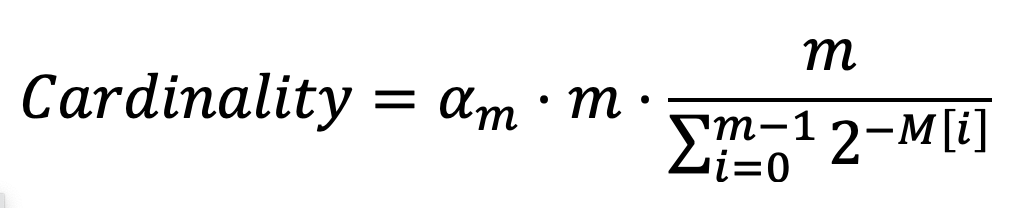
It is actually a normalized version of the harmonic mean of the 2M[i], where αm is a constant calculated by m as:
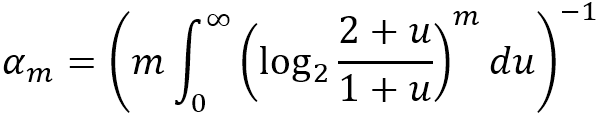
HyperANF
HyperANF is one popular ANF algorithm, it is a breakthrough improvement in terms of speed and scalability.
The algorithm is based on the observation that B(x,t), the set of reachable nodes from node x with distance t or less, satisfies
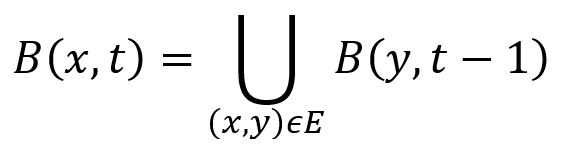
In the example graph below, node a has 3 adjacent edges (a,b), (a,c) and (a,d), so B(a,3) = B(b,2) ∪ B(c,2) ∪ B(d,2).
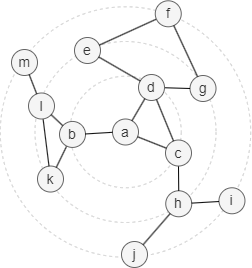
Instead of keeping tracking of B(x,t), the HyperANF algorithm employes HyperLogLog counters to simplify the computation process, as explained below with the above example graph:
- Each node x is mapped to a binary representation h(x), and is assigned a HyperLogLog counter Cx(t). Set b = 2, so each counter has m = 2b = 4 registers.
- Cx(0) is then computed by the value of i and ρ. Note: we use 0 instead of -∞ for the calculation, the result is the same.
- In the t-th iteration, for each node x, the union of
B(y,t-1)((x,y)∈E) is implemented by combining the counters of all neighbors of node x, that is, maximizing the values of the counter of node x register by register. - The value of all counters stay unchanged after 6 iterations, the reason is the diameter of the graph is 6.
|B(x,t)|is computed in each iteration by the cardinality equation with the constant αm = 0.53243.
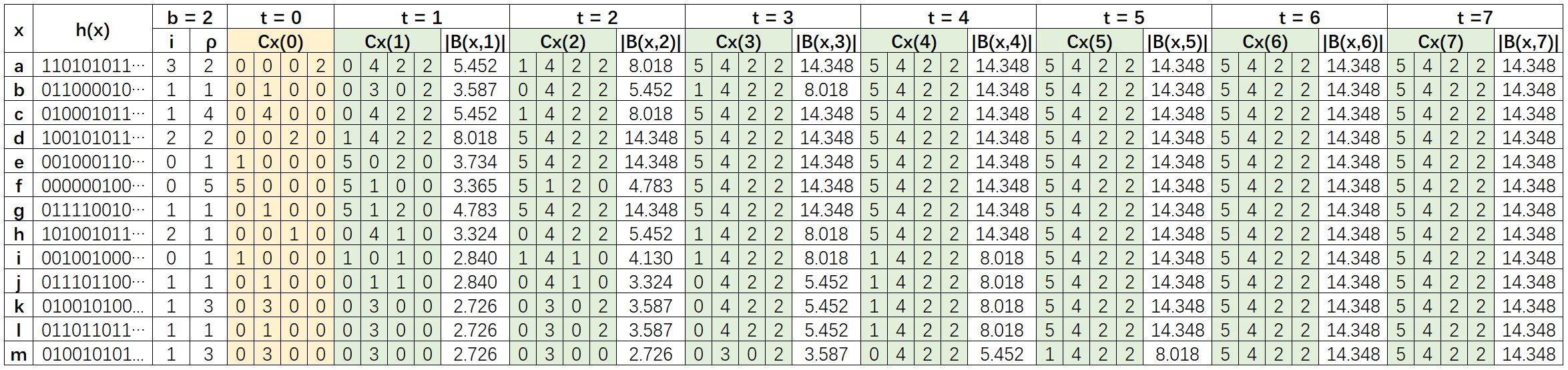
Since B(x,0) = {x}, then |N(x,t)| = |B(x,t)| - 1. In this example, the average graph distance computed by the algorithm is 3.2041. The exact average graph distance of this example is 3.
Considerations
- The HyperANF algorithm is typically best suited for connected graphs. For disconnected graphs, the algorithm may not provide accurate results.
- The HyperANF algorithm ignores the direction of edges but calculates them as undirected edges.
Example Graph
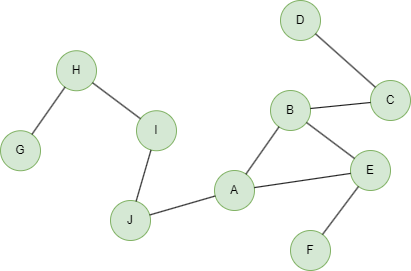
To create this graph:
// Runs each row separately in order in an empty graphset
insert().into(@default).nodes([{_id:"A"}, {_id:"B"}, {_id:"C"}, {_id:"D"}, {_id:"E"}, {_id:"F"}, {_id:"G"}, {_id:"H"}, {_id:"I"}, {_id:"J"}])
insert().into(@default).edges([{_from:"G", _to:"H"}, {_from:"H", _to:"I"}, {_from:"I", _to:"J"}, {_from:"J", _to:"A"}, {_from:"A", _to:"B"}, {_from:"A", _to:"E"}, {_from:"E", _to:"F"}, {_from:"B", _to:"E"}, {_from:"B", _to:"C"}, {_from:"C", _to:"D"}])
Creating HDC Graph
To load the entire graph to the HDC server hdc-server-1 as hdc_hyperANF:
CALL hdc.graph.create("hdc-server-1", "hdc_hyperANF", {
nodes: {"*": ["*"]},
edges: {"*": ["*"]},
direction: "undirected",
load_id: true,
update: "static",
query: "query",
default: false
})
hdc.graph.create("hdc_hyperANF", {
nodes: {"*": ["*"]},
edges: {"*": ["*"]},
direction: "undirected",
load_id: true,
update: "static",
query: "query",
default: false
}).to("hdc-server-1")
Parameters
Algorithm name: hyperANF
Name |
Type |
Spec |
Default |
Optional |
Description |
|---|---|---|---|---|---|
loop_num |
Integer | ≥1 | / | No | The maximum number of iteration rounds. The algorithm will terminate after completing all rounds. |
register_num |
Integer | [4,30] | / | No | The value of b which decides the number of registers (m = 2b) in the HyperLogLog counters. |
File Writeback
CALL algo.hyperANF.write("hdc_hyperANF", {
params: {
loop_num: 5,
register_num: 4
},
return_params: {
file: {
filename: "distance"
}
}
})
algo(hyperANF).params({
projection: "hdc_hyperANF",
loop_num: 5,
register_num: 4
}).write({
file: {
filename: "distance"
}
})
hyperANF_result: 2.46228
Stats Writeback
CALL algo.hyperANF.write("hdc_hyperANF", {
params: {
loop_num: 5,
register_num: 4
},
return_params: {
stats: {}
}
})
algo(hyperANF).params({
projection: "hdc_hyperANF",
loop_num: 5,
register_num: 4
}).write({
stats: {}
})
Result:
| hyperANF_result |
|---|
| 2.462277 |
Full Return
CALL algo.hyperANF("hdc_hyperANF", {
params: {
loop_num: 5,
register_num: 4
},
return_params: {}
}) YIELD distance
RETURN distance
exec{
algo(hyperANF).params({
loop_num: 5,
register_num: 4
}) as distance
return distance
} on hdc_hyperANF
Result:
| hyperANF_result |
|---|
| 2.462276 |
Stream Return
CALL algo.hyperANF("hdc_hyperANF", {
params: {
loop_num: 7,
register_num: 5
},
return_params: {
stream: {}
}
}) YIELD distance
RETURN round(distance.hyperANF_result)
exec{
algo(hyperANF).params({
loop_num: 7,
register_num: 10
}).stream() as distance
return round(distance.hyperANF_result)
} on hdc_hyperANF
Result:
| round(distance.hyperANF_result) |
|---|
| 3 |
Stats Return
CALL algo.hyperANF("hdc_hyperANF", {
params: {
loop_num: 7,
register_num: 10
},
return_params: {
stats: {}
}
}) YIELD distance
RETURN distance
exec{
algo(hyperANF).params({
loop_num: 7,
register_num: 10
}).stats() as result
return result
} on hdc_hyperANF
Result:
| bipartite_result |
|---|
| 2.903838 |

