Node2Vec
Overview
Node2Vec is a semi-supervised algorithm designed for feature learning of nodes in graphs while efficiently preserving their neighborhoods. It introduces a versatile search strategy that can explore both the BFS and DFS neighborhoods of nodes. It also extends the Skip-gram model to graphs for training node embeddings. Node2Vec was developed by A. Grover and J. Leskovec at Stanford University in 2016.
- A. Grover, J. Leskovec, node2vec: Scalable Feature Learning for Networks (2016)
Concepts
Node Similarity
Node2Vec learns a mapping of nodes into a low-dimensional vector space, intending to ensure that similar nodes in the network exhibit close embeddings in the vector space.
Nodes in network often shuttle between two kinds of similarities:
1. Homophily
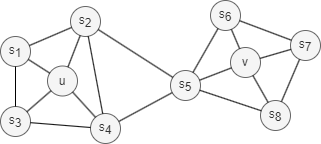
Homophily in networks refers to the phenomenon that nodes with similar properties, characteristics, or behaviors are more likely to be connected together or belong to the same or similar communities (nodes u and s1 in the graph above belong to the same community).
For example, in social networks, individuals with similar backgrounds, interests, or opinions are more likely to form connections.
2. Structural Equivalence
Structural equivalence in networks refers to the concept where nodes are considered equivalent based on their structural roles within the network. Nodes that are structurally equivalent have similar connectivity patterns and relationships to other nodes (i.e., the local topology), even if their individual characteristics are different (nodes u and v in the graph above act as hubs of their corresponding communities).
For example, in social networks, individuals that are structurally equivalent might occupy similar positions in their social groups.
Unlike homophily, structural equivalence does not emphasize connectivity; nodes could be far apart in the network and still have the same structural role.
When discussing structural equivalence, it's important to keep in mind two key points: Firstly, achieving complete structural equivalence in a real network is uncommon, leading us to focus on assessing structural similarity instead. Secondly, as the scope of the neighborhood being analyzed expands, the level of structural similarity between the two nodes tends to decrease.
Search Strategies
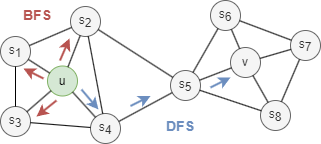
Generally, there are two extreme search strategies for generating a neighborhood set NS of k nodes:
- Breadth-first Search (BFS): NS is restricted to nodes which are immediate neighbors of the start node. E.g., NS(u) = s1, s2, s3 of size k = 3 in the graph above.
- Depth-first Search (DFS): NS consists of nodes sequentially searched at increasing distances from the start node. E.g., NS(u) = s4, s5, v of size k = 3 in the graph above.
The BFS and DFS strategies play a key role in producing embeddings that reflect homophily or structural equivalence between nodes:
- The neighborhoods sampled by BFS lead to embeddings that correspond closely to structural equivalence. By restricting search to nearby nodes, BFS obtains a microscopic view of the neighborhood which is often sufficient to characterize the local topology.
- The neighborhoods sampled by DFS lead to embeddings that correspond closely to homophily. By moving further away from the start node, DFS obtains a macro-view of the neighborhood which is essential in inferring node-to-node dependencies exist in a community.
Node2Vec Framework
1. Node2Vec Walk
Node2Vec employs a biased random walk with the return parameter p and in-out parameter q to guide the walk.
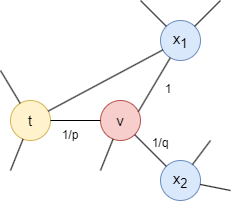
Consider the random walk that just traversed edge (t,v) and now arrives at node v, the next step of the walk is determined by the transition probabilities on edges (v,x) originating from v, which are proportional to the edge weights (weights are 1 in unweighted graphs). The weights of edges (v,x) are adjusted by p and q based on the shortest distance dtx between nodes t and x:
- If dtx = 0, the edge weight is scaled by
1/p. In the provided graph, dtt = 0. Parameterpinfluences the inclination to revisit the node just left. Whenp< 1, backtracking a step becomes more probable; whenp> 1, otherwise. - If dtx = 1, the edge weight remains unaltered. In the provided graph, dtx1 = 1.
- If dtx = 2, the edge weight is scaled by
1/q. In the provided graph, dtx2 = 2. Parameterqdetermines whether the walk moves inward (q> 1) or outward (q< 1).
Note that dtx must be one of {0, 1, 2}.
Through the two parameters, Node2Vec provides a way of controlling the trade-off between exploration and exploitation during random walk generation, which leads to representations obeying a spectrum of equivalences from homophily to structural equivalence.
2. Node Embeddings
The node sequences obtained from the random walks serve as input to the Skip-gram model. SGD is used to optimize the model's parameters based on the prediction error, and the model is optimized by techniques such as negative sampling and subsampling.
Considerations
- The Node2Vec algorithm ignores the direction of edges but calculates them as undirected edges.
Syntax
- Command:
algo(node2vec) - Parameters:
Name | Type | Spec | Default | Optional | Description |
|---|---|---|---|---|---|
| ids / uuids | []_id / []_uuid | / | / | Yes | ID/UUID of nodes to start random walks; start from all nodes if not set |
| walk_length | int | ≥1 | 1 | Yes | Depth of each walk, i.e., the number of nodes to visit |
| walk_num | int | ≥1 | 1 | Yes | Number of walks to perform for each specified node |
| edge_schema_property | []@<schema>?.<property> | Numeric type, must LTE | / | Yes | Edge property(-ies) to use as edge weight(s), where the values of multiple properties are summed up; nodes only walk along edges with the specified property(-ies) |
| p | float | >0 | 1 | Yes | The return parameter; a larger value reduces the probability of returning |
| q | float | >0 | 1 | Yes | The in-out parameter; it tends to walk at the same level when the value is greater than 1, otherwise it tends to walk far away |
| window_size | int | ≥1 | / | No | The maximum size of context |
| dimension | int | ≥2 | / | No | Dimensionality of the embeddings |
| loop_num | int | ≥1 | / | No | Number of training iterations |
| learning_rate | float | (0,1) | / | No | Learning rate used initially for training the model, which decreases after each training iteration until reaches min_learning_rate |
| min_learning_rate | float | (0,learning_rate) | / | No | Minimum threshold for the learning rate as it is gradually reduced during the training |
| neg_num | int | ≥0 | / | No | Number of negative samples to produce for each positive sample, it is suggested to set between 0 to 10 |
| resolution | int | ≥1 | 1 | Yes | The parameter used to enhance negative sampling efficiency; a higher value offers a better approximation to the original noise distribution; it is suggested to set as 10, 100, etc. |
| sub_sample_alpha | float | / | 0.001 | Yes | The factor affecting the probability of down-sampling frequent nodes; a higher value increases this probability; a value ≤0 means not to apply subsampling |
| min_frequency | int | / | / | No | Nodes that appear less times than this threshold in the training "corpus" will be excluded from the "vocabulary" and disregarded in the embedding training; a value ≤0 means to keep all nodes |
| limit | int | ≥-1 | -1 | Yes | Number of results to return, -1 to return all results |
Example
File Writeback
| Spec | Content |
|---|---|
| filename | _id,embedding_result |
UQLalgo(node2vec).params({ walk_length: 10, walk_num: 20, p: 0.5, q: 1000, buffer_size: 1000, window_size: 5, dimension: 20, loop_number: 10, learning_rate: 0.01, min_learning_rate: 0.0001, neg_number: 9, resolution: 100, sub_sample_alpha: 0.001, min_frequency: 3 }).write({ file:{ filename: 'embeddings' }})
Property Writeback
| Spec | Content | Write to | Data Type |
|---|---|---|---|
| property | embedding_result | Node Property | string |
UQLalgo(node2vec).params({ walk_length: 10, walk_num: 20, p: 0.5, q: 1000, buffer_size: 1000, window_size: 5, dimension: 20, loop_number: 10, learning_rate: 0.01, min_learning_rate: 0.0001, neg_number: 9, resolution: 100, sub_sample_alpha: 0.001, min_frequency: 3 }).write({ db:{ property: 'vector' }})
Direct Return
| Alias Ordinal | Type | Description | Columns |
|---|---|---|---|
| 0 | []perNode | Node and its embeddings | _uuid, embedding_result |
UQLalgo(node2vec).params({ walk_length: 10, walk_num: 20, p: 0.5, q: 1000, buffer_size: 1000, window_size: 5, dimension: 20, loop_number: 10, learning_rate: 0.01, min_learning_rate: 0.0001, neg_number: 9, resolution: 100, sub_sample_alpha: 0.001, min_frequency: 3 }) as embeddings return embeddings
Stream Return
| Alias Ordinal | Type | Description | Columns |
|---|---|---|---|
| 0 | []perNode | Node and its embeddings | _uuid, embedding_result |
UQLalgo(node2vec).params({ walk_length: 10, walk_num: 20, p: 0.5, q: 1000, buffer_size: 1000, window_size: 5, dimension: 20, loop_number: 10, learning_rate: 0.01, min_learning_rate: 0.0001, neg_number: 9, resolution: 100, sub_sample_alpha: 0.001, min_frequency: 3 }).stream() as embeddings return embeddings