Jaccard Similarity
Overview
Jaccard similarity, or Jaccard index, was proposed by Paul Jaccard in 1901. It’s a metric of similarity for two sets of data. In the graph, collecting the neighbors of a node into a set, two nodes are considered similar if their neighborhood sets are similar.
Jaccard similarity ranges from 0 to 1; 1 means that two sets are exactly the same, 0 means that the two sets do not have any element in common.
Concepts
Jaccard Similarity
Given two sets A and B, the Jaccard similarity between them is computed as:
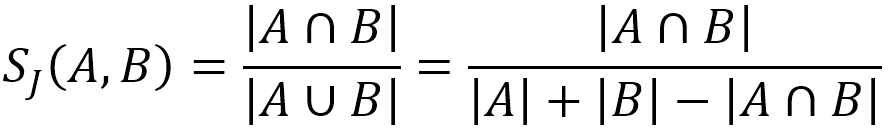
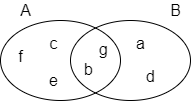
In the following example, set A = {b,c,e,f,g}, set B = {a,d,b,g}, their intersection A⋂B = {b,g}, their union A⋃B = {a,b,c,d,e,f,g}, hence the Jaccard similarity between A and B is 2 / 7 = 0.285714.
When applying Jaccard Similarity to compare two nodes in a graph, we use the 1-hop neighborhood set to represent each target node. The 1-hop neighborhood set:
- contains no repeated nodes;
- excludes the two target nodes.
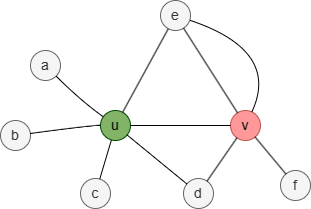
In this graph, the 1-hop neighborhood set of nodes u and v is:
- Nu = {a,b,c,d,e}
- Nv = {d,e,f}
Therefore, the Jaccard similarity between nodes u and v is 2 / 6 = 0.333333.
NOTEIn practice, you may need to convert some node properties into node schemas in order to calculate the similarity index that is based on common neighbors, just as the Jaccard Similarity. For instance, when considering the similarity between two applications, information like phone number, email, device IP, etc. of the application might have been stored as properties of @application node schema; they need to be designed as nodes and incorporated into the graph in order to be used for comparison.
Weighted Jaccard Similarity
The Weighted Jaccard Similarity is an extension of the classic Jaccard Similarity that takes into account the weights associated with elements in the sets being compared.
The formula for Weighted Jaccard Similarity is given by:
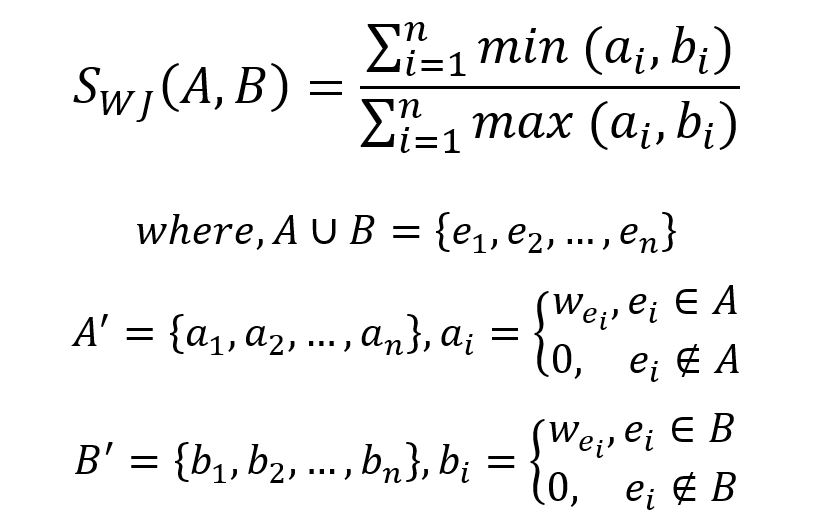
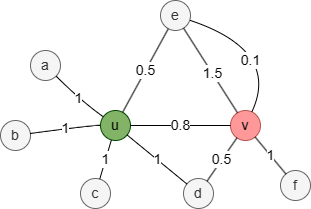
In this weighted graph, the union of the 1-hop neighborhood sets Nu and Nv is {a,b,c,d,e,f}. Set each element in the union set to the sum of the edge weights between the target node and the corresponding node, or 0 if there are no edges between them:
| a | b | c | d | e | f | |
|---|---|---|---|---|---|---|
| N'u | 1 | 1 | 1 | 1 | 0.5 | 0 |
| N'v | 0 | 0 | 0 | 0.5 | 1.5 + 0.1 =1.6 | 1 |
Therefore, the Weight Overlap Similarity between nodes u and v is (0+0+0+0.5+0.5+0) / (1+1+1+1+1.6+1) = 0.151515.
NOTEPlease ensure that the sum of the edge weights between the target node and the neighboring node is greater than or equal to 0.
Considerations
- The Jaccard Similarity algorithm ignores the direction of edges but calculates them as undirected edges.
- The Jaccard Similarity algorithm ignores any self-loop.
Syntax
- Command:
algo(similarity) - Parameters:
Name | Type | Spec | Default | Optional | Description |
|---|---|---|---|---|---|
| ids / uuids | []_id / []_uuid | / | / | No | ID/UUID of the first group of nodes to calculate |
| ids2 / uuids2 | []_id / []_uuid | / | / | Yes | ID/UUID of the second group of nodes to calculate |
| type | string | jaccard | cosine | No | Type of similarity; for Jaccard Similarity, keep it as jaccard |
| edge_weight_property | @<schema>?.<property> | Numeric type, must LTE | / | Yes | The edge property to use as edge weight, where the weights of multiple edges between two nodes are summed up |
| limit | int | ≥-1 | -1 | Yes | Number of results to return, -1 to return all results |
| top_limit | int | ≥-1 | -1 | Yes | In the selection mode, limit the maximum number of results returned for each node specified in ids/uuids, -1 to return all results with similarity > 0; in the pairing mode, this parameter is invalid |
The algorithm has two calculation modes:
- Pairing: when both
ids/uuidsandids2/uuids2are configured, pairing each node inids/uuidswith each node inids2/uuids2(ignore the same node) and computing pair-wise similarities. - Selection: when only
ids/uuidsis configured, for each target node in it, computing pair-wise similarities between it and all other nodes in the graph. The returned results include all or limited number of nodes that have similarity > 0 with the target node and is ordered by the descending similarity.
Examples
The example graph is as follows:
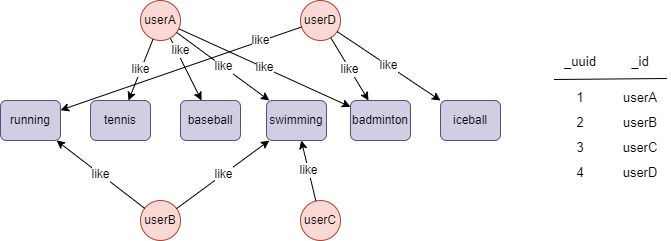
File Writeback
| Spec | Content |
|---|---|
| filename | node1,node2,similarity |
UQLalgo(similarity).params({ ids: 'userC', ids2: ['userA', 'userB', 'userD'], type: 'jaccard' }).write({ file:{ filename: 'sc' } })
Results: File sc
FileuserC,userA,0.25 userC,userB,0.5 userC,userD,0
UQLalgo(similarity).params({ uuids: [1,2,3,4], type: 'jaccard' }).write({ file:{ filename: 'list' } })
Results: File list
FileuserA,userC,0.25 userA,userB,0.2 userA,userD:0.166667 userB,userC:0.5 userB,userD,0.25 userB,userA,0.2 userC,userB,0.5 userC,userA,0.25 userD,userB:0.25 userD,userA:0.166667
Direct Return
Alias Ordinal | Type | Description | Columns |
|---|---|---|---|
| 0 | []perNodePair | Node pair and its similarity | node1, node2, similarity |
UQLalgo(similarity).params({ uuids: [1,2], uuids2: [2,3,4], type: 'jaccard' }) as jacc return jacc
Results: jacc
| node1 | node2 | similarity |
|---|---|---|
| 1 | 2 | 0.2 |
| 1 | 3 | 0.25 |
| 1 | 4 | 0.166666666666667 |
| 2 | 3 | 0.5 |
| 2 | 4 | 0.25 |
UQLalgo(similarity).params({ uuids: [1,2], type: 'jaccard', top_limit: 1 }) as top return top
Results: top
| node1 | node2 | similarity |
|---|---|---|
| 1 | 3 | 0.25 |
| 2 | 3 | 0.5 |
Stream Return
Alias Ordinal | Type | Description | Columns |
|---|---|---|---|
| 0 | []perNodePair | Node pair and its similarity | node1, node2, similarity |
UQLalgo(similarity).params({ uuids: [3], uuids2: [1,2,4], type: 'jaccard' }).stream() as jacc where jacc.similarity > 0 return jacc
Results: jacc
| node1 | node2 | similarity |
|---|---|---|
| 3 | 1 | 0.25 |
| 3 | 2 | 0.5 |
UQLalgo(similarity).params({ uuids: [1], type: 'jaccard', top_limit: 2 }).stream() as top return top
Results: top
| node1 | node2 | similarity |
|---|---|---|
| 1 | 3 | 0.25 |
| 1 | 2 | 0.2 |