Struc2Vec Walk
HDC
Overview
The Struc2Vec Walk is a biased random walk and serves as a key component of the Struc2Vec framework. Unlike traditional random walks, it operates on a constructed multi-layer weighted graph instead of the original graph. For more details, please refer to the Struc2Vec algorithm.
Example Graph
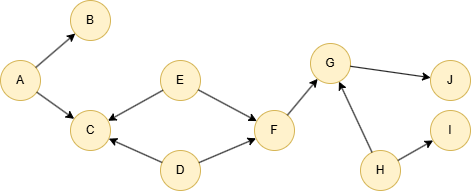
Run the following statements on an empty graph to define its structure and insert data:
INSERT (A:default {_id: "A"}), (B:default {_id: "B"}), (C:default {_id: "C"}), (D:default {_id: "D"}), (E:default {_id: "E"}), (F:default {_id: "F"}), (G:default {_id: "G"}), (H:default {_id: "H"}), (I:default {_id: "I"}), (J:default {_id: "J"}), (A)-[:default]->(B), (A)-[:default]->(C), (D)-[:default]->(C), (D)-[:default]->(F), (E)-[:default]->(C), (E)-[:default]->(F), (F)-[:default]->(G), (G)-[:default]->(J), (H)-[:default]->(G), (H)-[:default]->(I);
Creating HDC Graph
To load the entire graph to the HDC server hdc-server-1 as my_hdc_graph:
CREATE HDC GRAPH my_hdc_graph ON "hdc-server-1" OPTIONS { nodes: {"*": ["*"]}, edges: {"*": ["*"]}, direction: "undirected", load_id: true, update: "static" }
Parameters
Algorithm name: random_walk_struc2vec
Name | Type | Spec | Default | Optional | Description |
|---|---|---|---|---|---|
ids | []_id | / | / | Yes | Specifies nodes to start random walk by their _id. If unset, computation includes all nodes. |
uuids | []_uuid | / | / | Yes | Specifies nodes to start random walk by their _uuid. If unset, computation includes all nodes. |
walk_length | Integer | ≥1 | 1 | Yes | Depth of each walk, i.e., the number of nodes to visit. |
walk_num | Integer | ≥1 | 1 | Yes | Number of walks to perform for each specified node. |
k | Integer | [1, 10] | / | No | Number of layers in the constructed multilayer weighted graph, which should not exceed the diameter of the original graph. |
stay_probability | Float | (0,1] | / | No | The probability of walking in the current level. |
return_id_uuid | String | uuid, id, both | uuid | Yes | Includes _uuid, _id, or both values to represent nodes in the results. |
limit | Integer | ≥-1 | -1 | Yes | Limits the number of results returned. Set to -1 to include all results. |
File Writeback
algo(random_walk_struc2vec).params({ projection: "my_hdc_graph", return_id_uuid: "id", walk_length: 5, walk_num: 1, k: 4, stay_probability: 0.8 }).write({ file:{ filename: 'walks' }})
Result:
File: walks_ids J,G,F,E,C, D,F,E,F,E, F,G,F, H,I,H,F, B,A,B,A,B, A,C,E,D, E,F,G,H,G, C,D,F,G, I,H,G,F,E, G,D,F,
Full Return
exec{ algo(random_walk_struc2vec).params({ return_id_uuid: "id", ids: ['J'], walk_length: 6, walk_num: 3, k: 4, stay_probability: 0.8 }) as walks return walks } on my_hdc_graph
Result:
| _ids |
|---|
| ["J","G","F","C","B"] |
| ["J","F","H","F","J"] |
| ["J","G","J","H","F"] |
Stream Return
exec{ algo(random_walk_struc2vec).params({ return_id_uuid: "id", ids: ['J'], walk_length: 6, walk_num: 3, k: 5, stay_probability: 0.7 }).stream() as walks return walks } on my_hdc_graph
Result:
| _ids |
|---|
| ["J","G","I","F"] |
| ["J","E","J"] |
| ["J","H","F","A"] |