Preferential Attachment
Overview
Preferential attachment is a common phenomenon in complex networks, where nodes with more existing connections are more likely to attract new ones. When both nodes have a large number of connections, the probability of them forming a connection is significantly higher. This phenomenon was utilized by A. Barabási and R. Albert in their proposed BA model for generating random scale-free networks in 2002:
- R. Albert, A. Barabási, Statistical mechanics of complex networks (2001)
The Preferential Attachment algorithm measures the similarity between two nodes by multiplying the number of neighbors each node has. It is computed using the following formula:
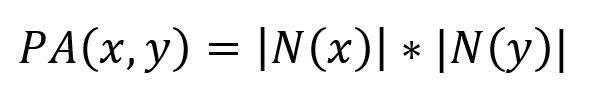
where N(x) and N(y) are the sets of adjacent nodes to nodes x and y respectively.
Higher Preferential Attachment scores indicate a greater similarity between two nodes, while a score of 0 indicates no such similarity.
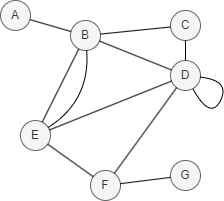
In this example, PA(D,E) = |N(D)| * |N(E)| = |{B, C, E, F}| * |{B, D, F}| = 4 * 3 = 12.
Considerations
- The Preferential Attachment algorithm treats all edges as undirected, ignoring their original direction.
Example Graph
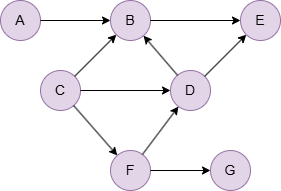
Run the following statements on an empty graph to define its structure and insert data:
INSERT (A:default {_id: "A"}), (B:default {_id: "B"}), (C:default {_id: "C"}), (D:default {_id: "D"}), (E:default {_id: "E"}), (F:default {_id: "F"}), (G:default {_id: "G"}), (A)-[:default]->(B), (B)-[:default]->(E), (C)-[:default]->(B), (C)-[:default]->(D), (C)-[:default]->(F), (D)-[:default]->(B), (D)-[:default]->(E), (F)-[:default]->(D), (F)-[:default]->(G);
Creating HDC Graph
To load the entire graph to the HDC server hdc-server-1 as my_hdc_graph:
CREATE HDC GRAPH my_hdc_graph ON "hdc-server-1" OPTIONS { nodes: {"*": ["*"]}, edges: {"*": ["*"]}, direction: "undirected", load_id: true, update: "static" }
Parameters
Algorithm name: topological_link_prediction
Name | Type | Spec | Default | Optional | Description |
|---|---|---|---|---|---|
ids | []_id | / | / | No | Specifies the first group of nodes for computation by their _id. If unset, all nodes in the graph are used as the first group of nodes. |
uuids | []_uuid | / | / | No | Specifies the first group of nodes for computation by their _uuid. If unset, all nodes in the graph are used as the first group of nodes. |
ids2 | []_id | / | / | No | Specifies the second group of nodes for computation by their _id. If unset, all nodes in the graph are used as the second group of nodes. |
uuids2 | []_uuid | / | / | No | Specifies the second group of nodes for computation by their _uuid. If unset, all nodes in the graph are used as the second group of nodes. |
type | String | Preferential_Attachment | Adamic_Adar | No | Specifies the similarity type; for Preferential Attachment, keep it as Preferential_Attachment. |
return_id_uuid | String | uuid, id, both | uuid | Yes | Includes _uuid, _id, or both to represent nodes in the results. |
limit | Integer | ≥-1 | -1 | Yes | Limits the number of results returned. Set to -1 to include all results. |
File Writeback
CALL algo.topological_link_prediction.write("my_hdc_graph", { ids: ["C"], ids2: ["A","E","G"], type: "Preferential_Attachment", return_id_uuid: "id" }, { file: { filename: "pa" } })
Result:
File: pa_id1,_id2,result C,A,3 C,E,6 C,G,3
Full Return
CALL algo.topological_link_prediction.run("my_hdc_graph", { ids: ["C"], ids2: ["A","C","E","G"], type: "Preferential_Attachment", return_id_uuid: "id" }) YIELD pa RETURN pa
Result:
| _id1 | _id2 | result |
|---|---|---|
| C | A | 3 |
| C | E | 6 |
| C | G | 3 |
Stream Return
CALL algo.topological_link_prediction.stream("my_hdc_graph", { ids: ["C"], ids2: ["A", "B", "D", "E", "F", "G"], type: "Preferential_Attachment", return_id_uuid: "id" }) YIELD pa FILTER pa.result >= 6 RETURN pa
Result:
| _id1 | _id2 | result |
|---|---|---|
| C | B | 12 |
| C | D | 12 |
| C | E | 6 |
| C | F | 9 |